CRISPR/Cas9 Knockout of Shell Matrix Protein 1 in the Slipper-Snail Crepidula atrasolea
Grant Batzel, Yiqun Wang, Antonia Bock, Elbereth Chen, Stephanie Neal, Rebecca N. Lopez-Anido, Yoon Lee, Evan Tjeerdema, Emily Ignatoff, Tejasvi Patil, Gabriela Ramirez, Maryna P. Lesoway, Amro Hamdoun and Deirdre C. Lyons
Journal of Experimental Zoology Part B: Molecular and Developmental Evolution
May 4, 2025
Abstract
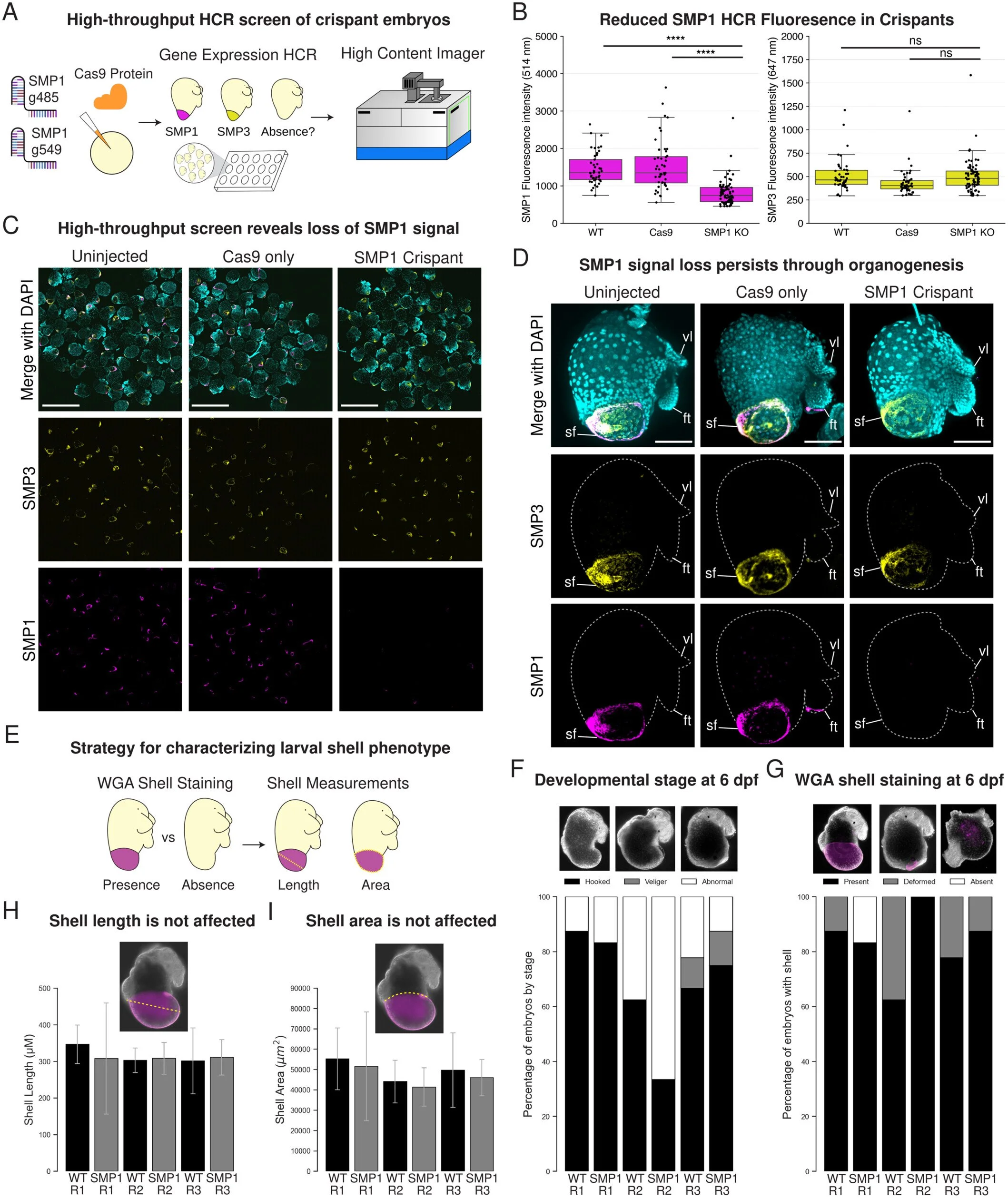
Over the course of hundreds of millions of years, biomineralization has evolved independently many times across all kingdoms of life. Among animals, the phylum Mollusca displays a remarkable diversity in biomineral structures, particularly the molluscan shell, which varies greatly in shape, size, pigmentation, and patterning. Shell matrix proteins (SMPs) are key components of these shells, and are thought to drive the precipitation of calcium carbonate minerals and influence shell morphology. However, this structure-function relationship has rarely been studied directly because tools for knocking out genes did not exist in molluscs until recently. In this study, we report the first successful use of CRISPR/Cas9 gene editing to target an SMP in gastropod molluscs. Using the emerging model gastropod Crepidula atrasolea, we generated knockouts of the SMP1 gene. Successful gene editing was confirmed by Sanger and MiSeq sequencing, and loss of SMP1 expression was validated through high-content imaging of crispant embryos. This study establishes C. atrasolea as a valuable model for investigating the genetic basis of shell formation and provides a framework for applying CRISPR/Cas9 technology in other molluscan species. Our approach will enable future studies to thoroughly test the role of SMPs in shaping the diverse array of molluscan shell structures.